Lecture: Data Quality Check and Cleaning
Actuarial Data Science - Open Learning Resource
Recommended Reading
- R for Data Science, Chapters 5, 7, 10
- Applied predictive Modelling, Chapters 3.3 (only transformations to resolve outliers), 3.4
Some coding examples in this lecture are adapted from R for Data Science (Wickham, Çetinkaya-Rundel, and Grolemund 2023).
Tibbles
Tibbles
- We work with “tibbles” instead of R’s traditional
data.framein thetidyverseenvironment. - The
tibblepackage provides opinionated data frames that make working in thetidyverseeasier. - See
vignette("tibble")for more information.
This lecture focuses on data quality: how to store your data in a tidy, convenient format, and how to identify and fix common data issues before they silently undermine your analysis. The aim is that by the end, you feel more confident trusting the datasets you work with, or at least knowing when you should not trust them yet.
Creating Tibbles
- Convert a data frame to a tibble:
as_tibble() - Create tibbles:
tibble() - Use non-syntactic names with backticks
tribble(): create tibbles in a transposed format
Example: Converting to a Tibble
Example: Creating a Tibble
Example: Creating Tibbles with Special Names
Pipe Data using %>%
- Use
%>%to emphasise a sequence of actions rather than the object being acted on - Pronounce
%>%as “then” when reading code - No need to create intermediate objects
%>%should always have a space before it and is usually followed by a new line.
Tibble vs. Data Frame
tibble()does less:- it never changes the type of the inputs (e.g. it does not convert strings to factors)
- it never changes variable names
- it never creates row names
- Printing
- tibbles show only the first 10 rows
- each column reports its type
- use
print()to display more rows (n) and columns (width)
- Subsetting
- extract a single variable using
$(by name) or[[ ]](by name or position) - within a pipe
%>%, use the special placeholder.
- extract a single variable using
Examples: Extracting Variables from a Tibble
[1] 0.06613743 0.64441979 0.87036981 0.99049084 0.76861869[1] 0.06613743 0.64441979 0.87036981 0.99049084 0.76861869[1] 0.06613743 0.64441979 0.87036981 0.99049084 0.76861869[1] 0.06613743 0.64441979 0.87036981 0.99049084 0.76861869[1] 0.06613743 0.64441979 0.87036981 0.99049084 0.76861869Import Data
Import Data
- Read Introduction to Data Science, Chapter 5
- Read R for Data Science, Chapter 7
Tidy Data
Tidy Data
- The same data can be organised in different ways.
- Tidy data is easier to work with.
- There are three interrelated rules that make a dataset tidy:
- Each variable has its own column
- Each observation has its own row
- Each value has its own cell
- Practical guidelines:
- Store each dataset as a tibble
- Store each variable in its own column
dplyr,ggplot2, and othertidyversepackages are designed to work with tidy data
Pivoting: Longer
- A common problem occurs when column names represent values rather than variables.
- Example:
table4a- Column names (1999, 2000) represent values of the
yearvariable - Values in these columns represent values of the
casesvariable - Each row represents two observations, not one
- Column names (1999, 2000) represent values of the
Pivoting: Wider
pivot_wider()is the opposite ofpivot_longer()- Use it when a single observation is spread across multiple rows.
- Example:
table2- Each observation is a country in a given year
- Data for each observation is spread across multiple rows
Separating
table3has a different problem: one column (rate) contains two variables (casesandpopulation).separate()splits one column into multiple columns, by splitting wherever a separator character appears.- By default,
separate()splits values at non-alphanumeric characters (i.e. characters that are not numbers or letters). - Use the
separgument to specify a custom separator.
Example: Using separate()
Unite
unite()is the inverse ofseparate(): it combines multiple columns into a single column- By default, it places an underscore (
_) between values from different columns - If no separator is desired, use
sep = ""
Missing Values
- A value can be missing in one of two ways:
- Explicitly: represented by
NA– the presence of an absence - Implicitly: not recorded in the dataset – the absence of a presence
- Explicitly: represented by
Making Implicit Missing Values Explicit
- Use
pivot_wider() - Use
complete()
Making Explicit Missing Values Implicit
- Set
values_drop_na = TRUEinpivot_longer()to convert explicit missing values into implicit ones
Filling Missing Values with fill()
fill()replaces missing values with the most recent non-missing value (also known as last observation carried forward).
Assessing Data Quality
Common Data Issues
- Missing data
- Irregular data and outliers
- Uninformative data
- Censored and truncated data
- High cardinality features
- Imbalanced data
Diagnosing Missing Data
- Descriptive statistics
- Visualisation (e.g. plots)
- See also: Missing values in tidy data; and Missing values in EDA
Missing Data
- Understand why values are missing
- Distinguish between structurally missing values and other types
- e.g. the number of children a man has given birth to
- Informative missingness: the pattern of missing data is related to the outcome
- e.g. in a drug study, side effects are so severe that patients drop out
Handling Missing Values
- Include a missing indicator (dummy variable)
- useful when the pattern of missingness is informative
- Some models can handle missing data (e.g. tree-based methods)
- Many models cannot handle missing values
- linear models, neural networks, SVMs
- Remove observations or variables as a last resort
- may be feasible for large dataset
- Impute missing values
Imputing Missing Values
- Use information from other predictors in the training data to estimate missing values
- simple methods: mean, median or mode
- model-based methods: e.g. k-nearest neighbours
- Imputation is extensively studied in the statistical literature for inference, but is often less critical in predictive modelling
Irregular Data and Outliers
- Detection:
- Descriptive statistics
- Visualisation (e.g. boxplots, scatter plots)
- Outlier detection methods
- Handling:
- Data validation: ensure there are no recording errors
- Remove or adjust values
- Outliers may belong to a different population
- Use models that are robust to outliers (e.g. tree-based methods, SVMs)
- Apply transformations to reduce the impact of outliers, such as the spatial sign transformation
- each observation is scaled by its norm
Example: Outliers
Uninformative Data
- Repetitive data
- Duplicate observations
- Irrelevant variables
- Collinearity: two or more predictor variables are highly correlated
- Redundant predictors increase model complexity
- Can lead to unstable models, numerical issues, and degraded predictive performance
- For linear regression, measures such as the variance inflation factor (VIF) can be used to identify predictors that are highly correlated with others
Censored data
- The value of an observation is only partially known
- For interpretation or inference
- Typically handled using formal methods with assumptions about the censoring mechanism
- For prediction
- Often treated as missing data or the censored value is used as observed
Examples:
General insurance (policy limits); life insurance (age groups of mortality data)
High-Cardinality Features
- Categorical predictors with many unique levels
- Examples: postcodes, medical condition codes, or similar variables
Some references on handling high-cardinality features:
- Dealing with features that have high cardinality
- Encoding High-Cardinality String Categorical Variables
- Similarity Encoding for Learning with Dirty Categorical Variables
- Nonlife Insurance Risk Classification Using Categorical Embedding
- Using Random Effects to Account for High-Cardinality Categorical Features and Repeated Measures in Deep Neural Networks
Imbalanced Data
- Imbalance between classes (e.g. control vs treatment) can cause modelling issues
- Construct a balanced training set to improve model performance
- Undersampling: reduce the number of observations in the majority class
- Oversampling: generate additional observations in the minority class
Data Validation
- Validate data against external sources and previous datasets
- Consult domain experts or data providers to assess data quality
Exploratory Data Analysis (EDA)
Exploratory Data Analysis
Generate questions about your data
Search for answers by visualising, transforming, and modelling your data
Use what you learn to refine your questions and/or generate new ones
We combine
dplyrandggplot2to interactively ask questions, answer them with data, and then ask new questionsTwo types of questions are especially useful for making discoveries in your data:
- What type of variation occurs within variables?
- What type of covariation occurs between variables?
Adapted from Wickham, Çetinkaya-Rundel, and Grolemund (2023), see Chapter 10 of R for Data Science
Variation
- Variation is the tendency of the values of a variable to change from measurement to measurement.
- Variables:
- Continuous variable: can take any value within a range of possible values
- Categorical variable: takes one of a fixed set of values
- Continuous variable: can take any value within a range of possible values
Visualising Distributios: Categorical Variables
- To examine the distribution of a categorical variable, use a bar chart
- Data:
diamonds(see?diamondsfor details)
See also: Covariation

Visualising Distributions: Continuous Variables
- To examine the distribution of a continuous variable, use a histogram

Exercise: Histogram Bin Widths
- Plot a histogram of diamonds with size less than 3 carats (using
filter()), and use a smaller binwidth of 0.1.

Exercise: Frequency Polygons by Group
- Overlay multiple histograms in the same plot by
cutusinggeom_freqpoly()

See also: Covariation
Typical Values
- In both bar charts and histograms, tall bars show common values of a variable, while shorter bars show less common values. Gaps indicate values that do not appear in the data.
- To turn this information into useful questions, look for anything unexpected:
- Which values are most common? Why?
- Which values are rare? Why? Does that match your expectations?
- Can you see any unusual patterns? What might explain them?
Example: Interpreting a Histogram
Question:
Look at the histogram below. What questions can you ask?

Example: Discussion
Why are there more diamonds at whole carats and common fractional values?
Why are there more diamonds just to the right of each peak than to the left?
Why are there no diamonds larger than 3 carats?
Unusual Values (Outliers)
- Outliers are observations that are unusual; data points that do not seem to fit the overall pattern
- Sometimes outliers are data entry errors; other times they may reveal important insights
- When you have a large dataset, outliers can be difficult to see in a histogram

Visualising Outliers
- To make unusual values easier to see, we can zoom in using
coord_cartesian(), often together withylim()orxlim(), to restrict the axis ranges.

Display Unusual Values
- Extract them using
dplyr. - What questions might you ask about these observations?
Deal with Outliers: Example
- In the ‘Diamond’ example, the
yvariable measures one of the three dimensions of a diamond (in mm) - Diamonds cannot have a width of 0 mm, so these values must be incorrect
- We might also suspect that measurements of 32mm and 59mm are implausible: such diamonds would be over an inch long, but do not cost hundreds of thousands of dollars
Deal with Outliers
- Repeat your analysis with and without the outliers.
- If they have minimal impact on the results and the cause is unclear, it is reasonable to replace them with missing values and proceed
- If they have a substantial impact on your results, do not remove them without justification
- Investigate the cause (e.g. data entry errors)
- Clearly document any decisions to remove them in your analysis
Missing Values
- If you encounter unusual values in your dataset and want to proceed with your analysis, you have two options:
- Drop the entire row containing the unusual values (not recommended — why?)
- Replace the unusual values with missing values (
NA) usingmutate()withifelse()orcase_when()
ggplot2does not include missing values in plots, but it will display a warning that they have been removed
Covariation
- If variation describes the behaviour within a variable, covariation describes behavior between variables.
- Covariation is the tendency for two or more variables to vary together in a related way
- The best way to identify covariation is to visualise relationships between variables.
- We consider three common cases:
- A categorical and a continuous variable
- Two categorical variables
- Two continuous variables
A Categorical and a Continuous Variable
- Explore the distribution of a continuous variable across levels of a categorical variable
- Use
geom_freqpoly()(e.g. see frequency polygon example) - It can be difficult to compare distributions when overall counts differ substantially (e.g. see distribution of cut example).
- Instead of displaying counts, we can display density, where the area under each curve is standardised to one

Boxplot
- Another way to display the distribution of a continuous variable across categories is a boxplot.
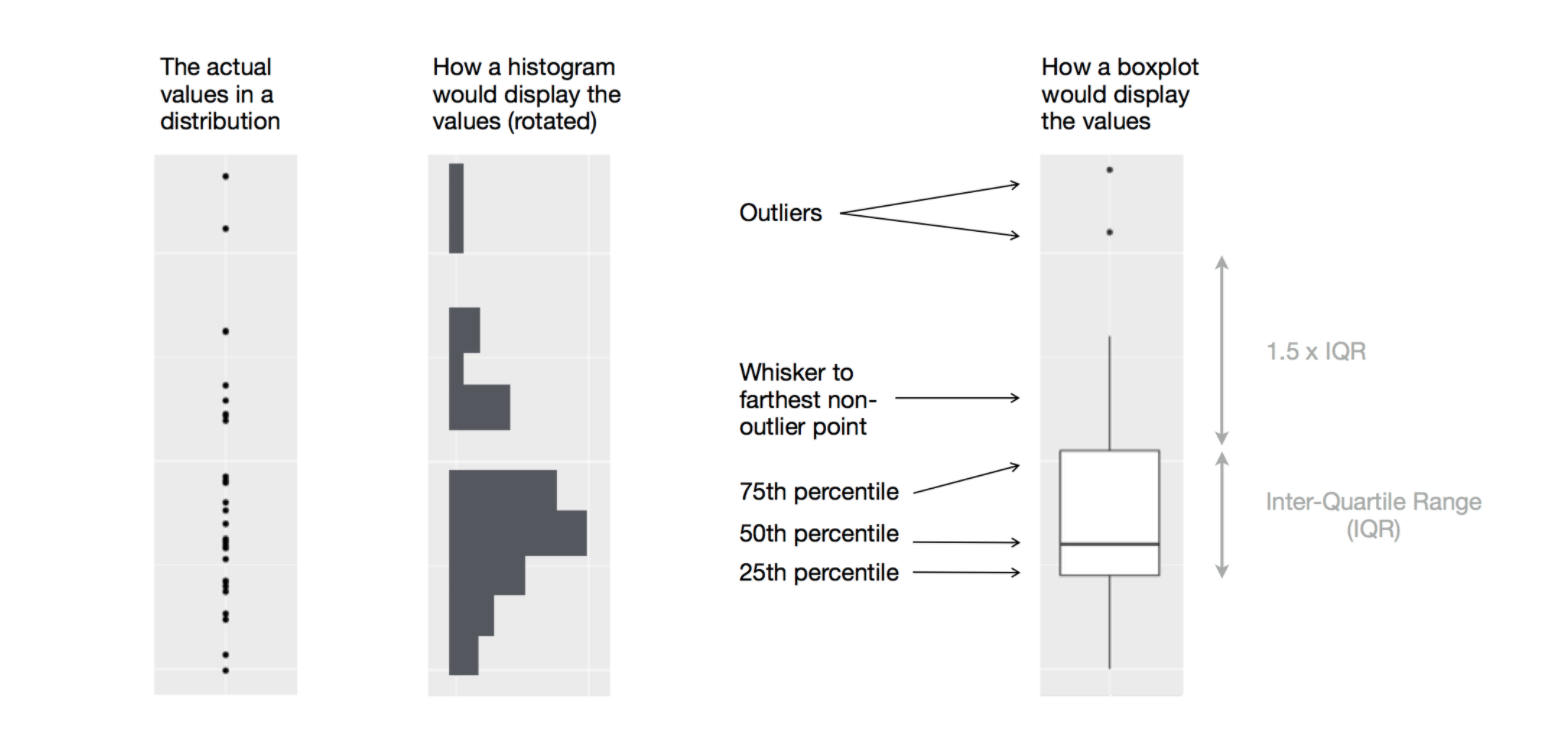
Diagram illustrating how a boxplot is constructed (Source: R for Data Science (Wickham, Çetinkaya-Rundel, and Grolemund 2023))
Example: Diamond Prices by Cut
- Examine the distribution of diamond prices by
cut. What do you observe?

- This suggests the counterintuitive finding that higher-quality diamonds are cheaper on average. Why might this be the case?
Example: Highway Mileage by Vehicle Class
- Consider the
mpgdataset. We want to understand how highway mileage (hwy) varies across vehicle classes (class) - To make patterns easier to see, reorder
classby the median ofhwyusingreorder(..., FUN = median) - For long variable names,
geom_boxplot()may be clearer when flipped usingcoord_flip()

Two Categorical Variables
- To visualise covariation between categorical variables, count the number of observations for each combination.
- One approach is to use
geom_count()

Example: Computing Counts with dplyr
- Another approach is to compute counts using
dplyr
Example: Visualising Counts with geom_tile()
- Then visualise the counts using
geom_tile()with the fill aesthetic

Two Continuous Variables
- A common way to visualise the covariation between two continuous variables is a scatter plot using
geom_point() - Covariation appears as patterns in the points
- Example: visualise the relationship between carat size and price of diamonds

Other Ways to Visualise the Relationship
- Use the
alphaaesthetic to add transparency - Use
geom_bin2d()orgeom_hex()to bin observations in two dimensions - Bin one continuous variable so that it behaves like a categorical variable
Example: Add Transparency

From Data Patterns to Models
- Patterns in your data provide clues about relationships
- Models are tools for extracting and formalising these patterns
References
